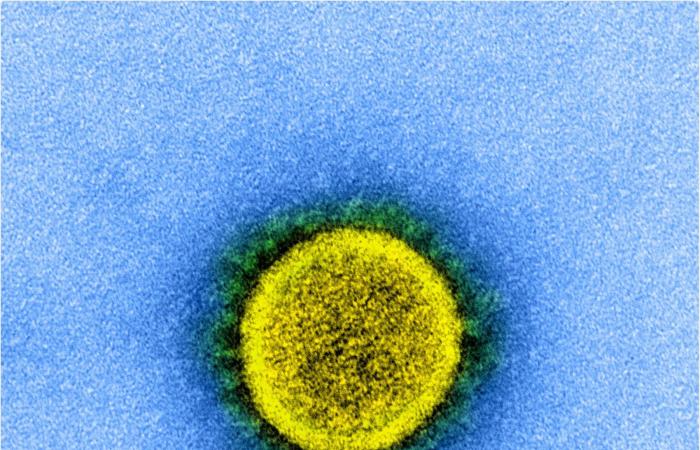
Transmission electron microscope image of a SARS-CoV-2 virus particle isolated from a patient. The image was captured and color enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Image Credit: NIAID
Die D614G-Mutation
The Wuhan variety, which spread throughout China and beyond to the rest of the world within a few weeks, has now spawned three main varieties. Among these, the Spike D614G mutation has been shown to offer a distinct advantage and is rapidly becoming dominant in every area of the world in which it was introduced. This mutation consists in replacing the aspartate residue at position 614 of the spike protein (or S protein) with a glycine residue.
The mutated variant of the spike seems to have increased the infectivity of the virus compared to that found in the wild type SARS-CoV-2. It is found to occur in two superclasses, namely S and G, but disease severity and lethality are only ever associated with one of them.
Currently, the S clade is predominant over much of Europe and Uruguay over the G clade, indicating that it is more contagious. The D614G mutation involves the substitution of a neutral amino acid, glycine, with a negatively charged one, aspartate. This is in the loop region of the S1 subunit of the spike protein.
This mutation is due to a single nucleotide variant in which A is replaced with G in the viral RNA. It didn’t appear in China until the last weeks of April 2020, but was then found in 90-97% of the samples. The mutation has made the virus more transmissible, with infections leading to higher viral loads. The question is whether this is linked to higher mortality, and whether this affects the effectiveness of vaccines by making the virus resistant to antibodies targeting the original strain of SARS-CoV-2.
In this case, the need for a revision could significantly delay the development of vaccines and other antivirals developed with the Wuhan reference strain in mind.
Differences in viral effects due to D614G
Some evidence from genetic studies, epidemiology and viral structural studies already show that this mutation is associated with a higher incidence of chemosensory deficits compared to the original strain. In addition, it shows a higher binding to the host cell receptor, the angiotensin converting enzyme 2 (ACE2), which enables a more efficient virus entry due to the higher stability of the spike protein and the reduced cleavage.
Genetic diversity and polymorphism of SARS-CoV-2
The current study aimed to examine the genetic diversity and polymorphisms of the viral spike protein. The researchers found that out of all 18 sequences derived from Malaysian and Venezuelan sequences uploaded to the National Center for Biotechnology Information (NCBI), there were 57 polymorphic sites among nearly 30,000 base pairs analyzed. These were parsimony informative sites, with at least two nucleotides showing a variation and at least two of the nucleotide variants showing a frequency of two or more. Further examination of these sites revealed the existence of two different groups, and the researchers suspect that these may share haplotypes in both countries.
Two genetic variants that show a different development
The researchers found that there were marked differences within the population, and an analysis of the fixation index (FST score) confirmed the presence of two different genetic variants with clearly observed variations both within and between the two groups. However, all Malaysian tribes were genetically very similar, while the Venezuelan tribes showed greater evolutionary divergence.
All haplotypes, the sets of genes inherited together from a parent, showed that mutation in the form of transversions and transitions was actively taking place. Insertion-deletion (Indel) mutations were only found within haplotype groups in the Venezuelan samples. Both the Tajima and Fs de Fu tests showed that there was no population growth within the Malaysian group. The mutation rates were stable between different locations.
The time of evolutionary divergence was observed in all populations using all previous methods. However, the researchers summarized the structuring of the two groups as being slightly higher in the Venezuelan group, possibly due to genetic flow.
Consistent results from different methods indicate conservation
The researchers comment that the discontinuous pattern of genetic divergence between groups “supports the idea of possible isolations due to past fragmentation events,” especially since the number of branches in the tree is small and few steps are required to be created.
The few mutations that have occurred appear to have escaped correction by genetic drift or have not experienced the basic effect that typically occurs when haplotypes are dispersed or lost over time. Taking genetic distance into account, the researchers found that the groups still didn’t show much difference. The researchers think it is important to take into account the minimal differences between the two groups from Malaysia and Venezuela when exchanging haplotypes.
The researchers found from their dendrogram that the main differences in the strains of the virus in both countries were related to their shape rather than their number. They concluded that the 18 strains in the two geographic regions could be distinguished due to the significant differences in the haplotypes, showing a hierarchical structure for all components of covariance. and also by the differences within and between the two groups. The estimated mean divergence between these tribes and other countries is absent or at least minimal.
Because many haplotypes were common in the Venezuelan group, the researchers did not attempt to find the link between genetic distance and geographical separation. The use of the estimator φ, an indicator that responds to the smallest molecular variation, gives credibility to the results of the various tests used.
You say this appraiser “supported the consistency between the results of all methods used and can be interpreted as phylogenetic confirmation that there is a consensus on the maintenance of the mutated SARS-CoV-2 genome (D614G) in the two countries analyzed. “
Implications
The implications are that the small number of polymorphisms currently found are unlikely to represent large changes in the proteins expressed by different strains of the virus. Therefore, despite the differences in haplotypes in these two geographic regions, these are not significant. Therefore, it makes sense to transfer these results to other variants in other countries and to conclude that they do not reflect any significant differences in the protein products of the wild type and the mutated SARS-CoV-2.
The study comes to the conclusion: “The analyzes conducted in this study suggest small variations in protein products, especially among those targeted by this study, and bring peace of mind about the risk of death from COVID-19, as well as potential problems adjusting some molecular targets for vaccines. ”
* Important NOTE
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and should therefore not be considered conclusive, guide clinical practice / health-related behavior, or are treated as established information.
These were the details of the news An analysis of the genetic diversity of the D614G mutation in... for this day. We hope that we have succeeded by giving you the full details and information. To follow all our news, you can subscribe to the alerts system or to one of our different systems to provide you with all that is new.
It is also worth noting that the original news has been published and is available at de24.news and the editorial team at AlKhaleej Today has confirmed it and it has been modified, and it may have been completely transferred or quoted from it and you can read and follow this news from its main source.