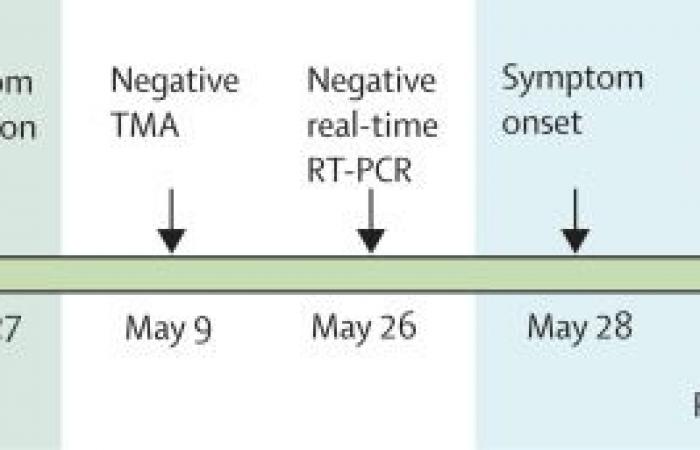
Procedure
The samples were taken from the patient with a nasopharyngeal swab at the group examination, during the isolation and recovery period, and when presented to the hospital. The swabs were transported to the Nevada State Public Health Laboratory (Reno, NV, USA) either with viral transport medium or with Aptima Multiswab Transport Media (Hologic, San Diego, CA, USA). The samples were transported on cold packs and stored refrigerated (4-8 ° C) for no longer than 72 hours before nucleic acid extraction and subsequent real-time RT-PCR.
Nucleic acid extraction was carried out using the Omega Biotek MagBind kit for viral DNA / RNA 96 (Omega Bio-tek, Norcross, GA, USA) according to the manufacturer’s instructions and with an elution volume of 100 μl. l carried out. Aliquots of eluted RNA were subjected to real-time RT-PCR, either using the Taqpath COVID-19 Multiplex Assay (ThermoScientific, Waltham, MA, USA; 10 μL aliquots) or the US Centers for Disease Control and Prevention (CDC)) 2019-nCoV real-time RT-qPCR diagnostic panel (CDC, Atlanta, GA, USA; 5 μL aliquots). Samples transported on Aptima Multiswab Transport Media were tested by transcription-mediated amplification using the Aptima SARS-CoV-2 assay (Panther System) (Hologic, Marlborough, MA, USA). The tests were performed according to the respective EUA procedures, unless otherwise stated. For the Taqpath real-time RT-PCR test, the sample recall threshold was positive for the reactivity of two out of three target sequences, each with a reactivity at a cycle threshold of less than 40.00. A positive or negative result from the Hologic Aptima assay was based on proprietary processes. The antibody test was carried out with the Roche Elecsys Anti-SARS-CoV-2 test (Roche Diagnostics, Indianapolis, IN, USA).
For viral genome sequencing, the total RNA was extracted from nasopharyngeal swabs as described. 70 µ l extracted RNA were treated with Qiagen DNase I (Qiagen, Germantown, MD, USA) for 30 min at room temperature and then purified and purified with silica spin columns (Qiagen RNeasy MinElute; Qiagen) with a 12. l concentrated water elution. A portion (7 μl) of this RNA was bound to an rRNA inhibitor (Qiagen FastSelect rRNA HMR; Qiagen) and then reverse transcribed (cDNA) using random hexamers. The synthesized DNA was strand-ligated and amplified isothermally to form micrograms of DNA (Qiagen FX Single Cell RNA Library Kit; Qiagen). A portion (1 µg) of this amplified DNA was sheared and ligated to Illumina-compatible sequencing adapters, followed by six cycles of PCR amplification (KAPA HiFi HotStart library amplification kit; Roche Sequencing and Life Science, Kapa Biosystems, Wilmington, MA, USA) ) for library molecules with adapters at both ends. Next, these sequencing libraries were enriched to a sequence specific for SARS-CoV-2 using biotinylated oligonucleotide baits (myBaits Expert Virus, Arbor Biosciences, Ann Arbor, MI, USA). A further eight to 16 PCR cycles were carried out after enrichment (98 ° C for 45 s, 98 ° C for 15 s, 60 ° C for 30 s, repetition for eight to 16 cycles, then 72 ° C for 60 s and 4 ° C), and these SARS-CoV-2 -enriched sequencing libraries were pooled and sequenced with an Illumina NextSeq 500 (Illumina, San Diego, CA USA) as paired 2×75 base pair reads using the NextSeq version 2.5 Medium power 150 stroke kit (Illumina).
For the bioinformatic analysis of the two SARS-CoV-2 agents (here referred to as sample A and sample B), after sequencing each library, FASTQ files were transferred to the CLC Genomics Workbench Version 20.0.4 (Qiagen A / S, Vedbæk, Denmark) with the CLC Microbial Genomics Module, the CLC Genome Finishing Module and the Biomedical Genomics Analysis. In short, the readings were imported, cropped and assigned to the SARS-CoV-2 reference sequence MN908947.3 of the National Center for Biotechnology Information. The alignment was refined with the help of the InDels and Structural Variants module, followed by the Local Realignment module. Variants were identified with a minimum coverage of five reads, a minimum number of five, and a minimum frequency of 70.0%.
Fast lift-read alignment with bowtie 2.
Optical PCR duplicates were marked with Picard MarkDuplicates in Picard-Slim version 2.22.5. Variants were called for both samples along with Freebayes version 1.0.2 with ploidy settings of 1, a minimum allele frequency of 0.70, and a minimum depth of five reads for each variant call. The genomic sequence of each sample was constructed using coverage statistics from BBtools stackup.sh and applyvariants.sh version 38.86, with only variants supported by coverage of five or more reads written to the bcftools consensus version 1.10.2 and all positions supported by less than five reads, whether reference or alternative, were replaced with Ns.
A statistical framework for SNP calls, mutation detection, association mapping and population genetic parameter estimation from sequencing data.
- Hartley P.
- Tillett RL
- Xu Y.
- et al.
Genomic monitoring revealed the prevalence of unique SARS-CoV-2 variants that carry a mutation in the RdRp gene in Nevada patients.
the SARS-CoV-2 reference strain (MN908947.3) and a sequence derived from isolate USA-WA1 / 2020 (Bei Resources, Manassas, VA, USA). After trimming six 5 ‘bases (Ns) from Sample A and 98 Ns from Sample B, the genomic sequences were aligned and related using NGPhylogeny.fr PhyML + SMS.
- Lemoine F.
- Correia D.
- Lefort V.
- et al.
NGPhylogeny.fr: New generation phylogenetic services for lay people.
The sequences were then first aligned using MAFFT with automatic flavor selection.
MAFFT Multiple Sequence Alignment Software Version 7: Improvements in performance and usability.
Informative regions were selected using block mapping and gathering with entropy, a sliding window size of 3 and a maximum entropy of 0.5.
BMGE (Block Mapping and Gathering with Entropy): A new software for the selection of phylogenetically informative regions from several sequence alignments.
Unrooted trees were created by PhyML using Smart Model Selection, the Akaike Information Criterion, and Subtree Pruning and Regrafting.
- Lefort V.
- Longueville I.
- Gascuel O.
SMS: Smart Modellauswahl in PhyML.
Newick trees were visualized with Interactive Tree Of Live version 4 and rooted on the Wuhan reference stem.
Interactive Tree of Life (iTOL) v4: Current updates and new developments.
Important SARS-CoV-2-Clade memberships were predicted with Nextclade.
National Institute for Standards and Technology
Population studies by the NIST Forensics / Human Identity Project team.
These were the details of the news Genomic evidence for reinfection with SARS-CoV-2: a case study for this day. We hope that we have succeeded by giving you the full details and information. To follow all our news, you can subscribe to the alerts system or to one of our different systems to provide you with all that is new.
It is also worth noting that the original news has been published and is available at de24.news and the editorial team at AlKhaleej Today has confirmed it and it has been modified, and it may have been completely transferred or quoted from it and you can read and follow this news from its main source.